Rädler Group
We are a soft matter physics group studying self-assembly and dynamics of macromolecules and lipid membranes and the spatio-temporal self-organization of living cells (active matter).
We are a soft matter physics group studying self-assembly and dynamics of macromolecules and lipid membranes and the spatio-temporal self-organization of living cells (active matter).

Join the Rädler Group's weekly seminar featuring invited speakers, primarily from the life sciences, sharing insights into cutting-edge research. 🗓️ Wednesdays, 4 p.m. during the lecture period.
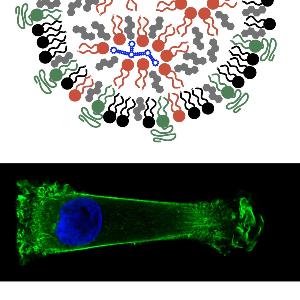
In gene and siRNA based therapy, therapeutic benefits rely on the efficient transfer of foreign nucleic acids into cells. We investigate synthetic complexes capable of readily carrying nucleic acid across the cell membrane. The self-assembly, size and structure of lipid- and polycation-DNA complexes, known respectively as lipoplexes and polyplexes, are studied. We also endeavor to quantify and model transfection at the single cell level using GFP reporter and single cell time-lapse microscopy.
Read more
Collective cell migration is a key element in morphogenesis, cancer formation and wound healing. The multicellular dynamics of epithelial cell layers were studied in experiment and theory over the last years. Yet little is known about cell patches of finite size that should lead over from single cell behavior to collective motion. Here we study the migration and proliferation of small assemblies of epithelial cells.
Read more
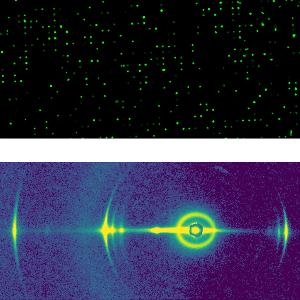
We are interested in the kinetics of cellular responses, in particular in gene expression after transfection and event cascades in cytotoxicity. Using live cell imaging on single cell arrays (LISCA) time courses from individual cells are monitored and analyzed based on mathematical models. Cellular microarrays help to resolve cellular heterogeneity, a hallmark of complex biological systems dominated by intrinsic fluctuations.
Read more
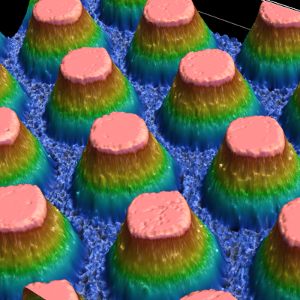

We fabricate artificially designed micro-environments to culture and probe living cells. Arrays of defined cavities provide standardized geometric boundaries for individual cells or groups of cells. We study single cell gene expression, cell migration and apoptotic events with precise spatio-temporal control. To this end we develop biofabrication techniques, including photo-lithography, hot-embossing, electroplating, soft lithography, 3D printing and microcontact printing, to create two-dimensional (2D) and three-dimensional (3D) pattern, cavities and channels made of biocompatible materials like PDMS, COC, hydrogels and proteins. The 3D microenvironments allow for long-term observation of growth and function of individual living cells, interacting groups of cells and organoids.
Supported membranes are versatile biomimetic interfaces. We study the structure and dynamics of supported membranes on various solid supports in collaboration with the group of Bert Nickel. The control of fluidity, spacing and lateral organisation is a current research challenge.
Find a list of all publications by the Rädler Group on this page.
If you would like to do your bachelor or master thesis in our group, or to apply for a PhD position, please contact Prof. Joachim Rädler.
We sometimes post offers for specific projects, which you can find on the Open Positions page, or below:
Living tissues are active materials that have the ability to self-organize and repair themselves. Their macroscopic properties, such as rigidity and fluidity, emerge from the properties and interactions of their constituent cellular units. In the case of cancer, diseased cells coexist with healthy cells as a two-component cell mixture. Altered cell deformability and cell–cell interactions result in segregation, domain formation and metastasis.
In the Rädler group, we study the multicellular dynamics of epithelial cells by using a minimal system of four cells confined in micropattern. We have recently developed a method to induce repeated T1-transitions of the cell junctions, enabling reproducible measurements of cell-cell decoupling.
As a Master’s student, your task will be to take fluorescence life cell time-lapse microscopy data and to quantitatively derive cell interaction parameters using state-of-the art image analysis and mathematical modeling. The project will be carried out in collaboration with the dynamics of living systems group of Wolfgang Losert at the University of Maryland (USA) including student exchange opportunities.
If you are interested and would like to learn more, please contact Prof. Dr. Joachim Rädler or Agathe Jouneau.
| Name | Position |
|---|---|
| Rädler, Joachim | |
| Meixner, Margarete | Administration |
| Leu, Charlott | Technical Staff |
| Schwake, Gerlinde | Technical Staff |
| Rüter-Stimpfle, Margarita | Administration |
| Jouneau, Agathe | PhD Student |
| Philipp, Julian | PhD Student |
| Maksutova, Daria | PhD Student |
| Stockbauer, Simon | Master student |
| Kostyurina, Ekaterina | Postdoc |
| Kirchmair, Bernhard | PhD Student |
| Kuhl, Santiago | PhD Student |